AI & Sci roundup #2: More bio data & better models? Or something else?
Thinking about biology data in layers, wishing for APIs in drug development, and celebrating those who test their models in the lab!
Hello all! It’s SynBioBeta week and I am finally going! It has been years I’ve been eyeing this meeting. This year I see it less as just synthetic biology and more as an AI x bio conference. I’m sure I’m not the only one.
In light of this I’ve been reading a lot of exciting posts and seeing lots of cool events happening. I wanted to take the time to round up some of what I’ve been reading and enjoying!
Pick #1: On biological data generation (more is different, and so is the data)
First, this post by Michael Koeris at DARPA. Mike is a longtime ‘phage friend’ of mine, as before DARPA he started two phage companies (and was kindly one of the first to pay attention to what we were doing at Phage Directory back in ~2018!). He was also on the board of Addgene, one of the best science nonprofits I know of.
In his post he talks about machine learning-ready bio, one of my favourite topics these days. What makes biology data good for machine learning? He gets into why virtual cell needs to be made of a lot more kinds of data, and describes really nicely why we should think of it in layers — the cell is a layer, but so is the sub-cell (organelles), molecules, and then of course zooming out we have tissues, organs, beings. There needs to be data collected at all these different layers, though not necessarily all at once. It depends on the question you’re asking.
He also describes cells as things with lots going on inside, but only a subset of that stuff being presented to the outside world (ie. to other cells). If we are building models just on what’s going on inside cells, we’re probably not developing a true model of the system, since not everything inside is actually presented to the outside. If we’re wanting to model tissues, cell-cell interactions matter, so that layer requires looking at what cells are actually presenting to each other (ie. what receptors? what secreted signals?).
I thought this was a refreshing take, and gave me some language and visuals (if only in my mind) to think about what ‘ML ready bio’ could mean. I also enjoyed that it wasn’t just the ‘virtual cell is dumb / too ambitious’ take I’ve been seeing go around X lately.
If this interests you, check out Mike’s post here:
And then if you’re keen to learn even more, go find Mike at SynBioBeta this week (it’s not too late to register!), where he’ll be speaking at 9 am on Wednesday on a biosecurity panel, at 3:30 pm the same day in a breakout session (this one is on AI-ready bio data factories, so this would be the one most relevant to the post). And at 4:30 pm that day you can also go to his session on how to get government money if you’re a founder.
Pick #2: AI for bio has a fuzzy API problem
Another great post I came across this week was by Ankit Gupta, General Partner at Y Combinator. He talks about why AI for bio is hard, and gets at why it’s not just because it’s complex (since AI/ML is indeed great at lots of complex things). It’s because the components aren’t well defined and the feedback loops aren’t fast. There aren’t clear handoff points between well-defined components, so it becomes hard for companies to specialize in just one component and hand that off to the next. Where the computer world has APIs (well-defined ways a component of a system can be reliably communicated with), the biological/drug development world does not. We abstract a lot of steps into one (ie. target discovery is actually a slew of things, many of which are hypotheses; clinical trials are a slew of a bunch of other things). As a result, the output of one step isn’t always what you expect (a target isn’t always a good target; a clinical trial could be done perfectly and still not work because it was done on the wrong kinds of patients).
So because we don’t have the system defined into chunks where each have defined inputs/outputs, it’s hard to train models on it. So Ankit’s point is that many people starting companies (often coming from computer science backgrounds where there ARE really well established APIs and ways to know if they’re broken right away) focus on narrow chunks of the puzzle and expect that to be enough. Like making the next Boltz model. But doing a great job in one area, when the rest of the system isn’t ready for your thing to be plugged into, will likely not be as successful as you’d hope.
Ankit advocates for more vertically integrated approaches, like using AI to find underevaluated drug modalities, and going to China to license a drug candidate that purports to do that, and doing the clinical development yourself. Or collecting a new kind of data that lets you come up with new drug targets, getting good at rapidly testing them, and then licensing those candidates to pharma. Both approaches still use AI/ML, and are pointed at drug development, but aren’t so focused on just one layer, expecting the rest of the system to plug into it, like software companies can plug into Stripe.
Check out Ankit’s post here if you want to learn more!
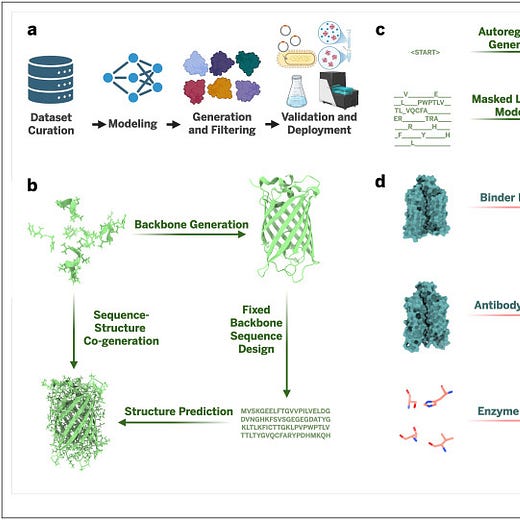
Pick #3: Closing the loop: Experimentally validated methods in AI–driven protein design
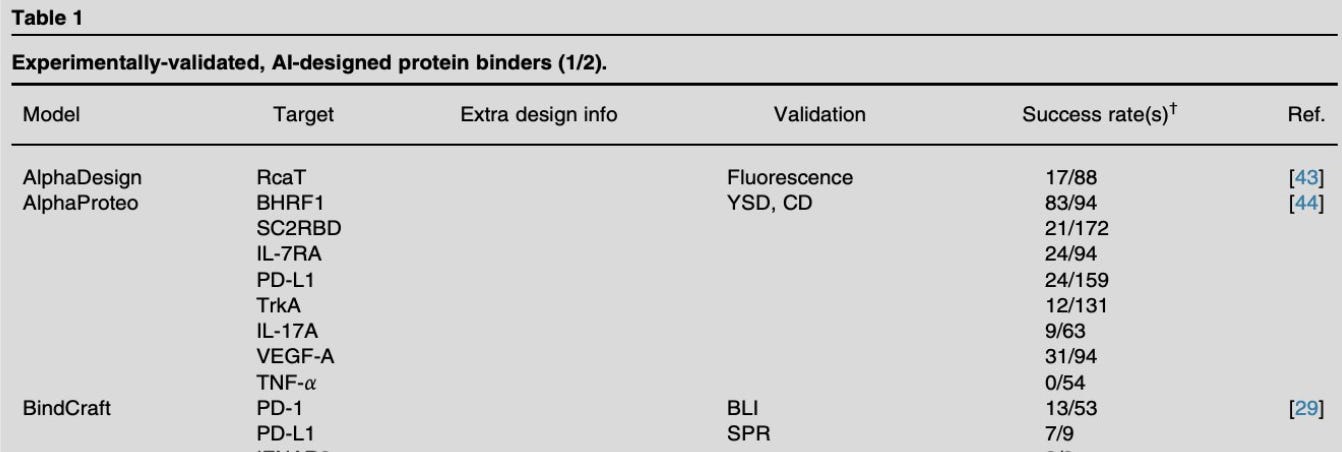
This last pick is one I want to read but haven’t yet. It’s a review paper where they compiled and compared the wet lab success rates of 200+ different protein design tasks.
One of the authors, Clay Kosonocky, posted this nice thread as well (this is how I came across it):
I bookmarked it because I really want to get my head around how the wet lab best connects with AI/ML models being developed. I realized in seeing his tweet that I wasn’t really sure what the typical success rates are these days for models tested in the lab.
One paper I refer to all the time, this one by Samuel King and colleagues at Arc Institute, looks at AI-designed phages (my brain can more easily understand it because phages are what I study). In it, they generated about 4000 designs, chose about 300 of them to test in the lab, and about 16 of them ‘worked’ (in this case ‘worked’ means the AI-generated phage genome sequences successfully came to life and killed bacteria). So this ‘success rate’ was in my head as what one might encounter. Is this good? I don’t know. For the phage field, it’s the state of the art! No one else is doing this. But what is a good success rate in the lab for other more popularly worked-on AI-generated things?
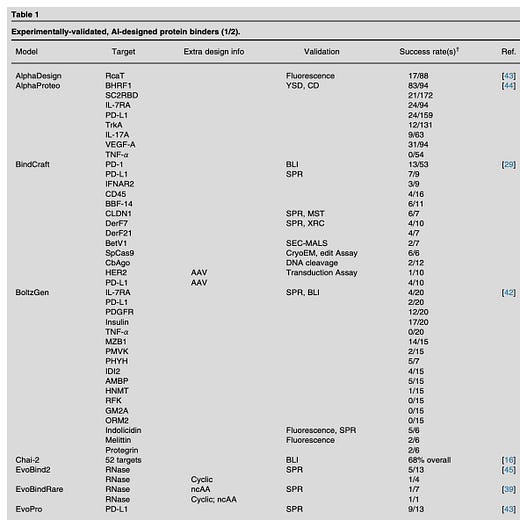
So this paper seems to get at that answer. You can see from their table that they’ve looked at a bunch of papers, and for different models, many targets have been tested against, with different validation methods (assays). The success rates are there: some high, some low. Mostly I’m noticing the denominator (number tested in the lab, I assume) is in the 2-3 digit range (tens to low hundreds being tested). The percent success is across the board at first glance; especially interesting to see one model tested on multiple targets gave quite different success rates.
Anyway there’s obviously a ton to dive into in Clay’s paper, so check it out if you’re interested in this too!
Mostly I just want to highlight that I think this is a great use of time, going into the literature to actually pull this kind of data out of a bunch of papers, so we the readers can get a sense of what the ‘going rate’ of model success rates is. And for everyone to see that the the wet lab is an important component of giving feedback to the models being developed (of course! I don’t think anyone is really arguing with this). It’s also just nice to see the analysis and bookmark it as a reference, so when you see a new tweet about a model and how great it is, you’re prompted to think ‘did they test it in the lab? How successful was it compared to [competing model xyz']?
Obviously a paper is going to be out of date quickly; it would be cool if there was an updating dashboard on this stuff! If anyone is aware of one, I’d love to know!
Events this week!
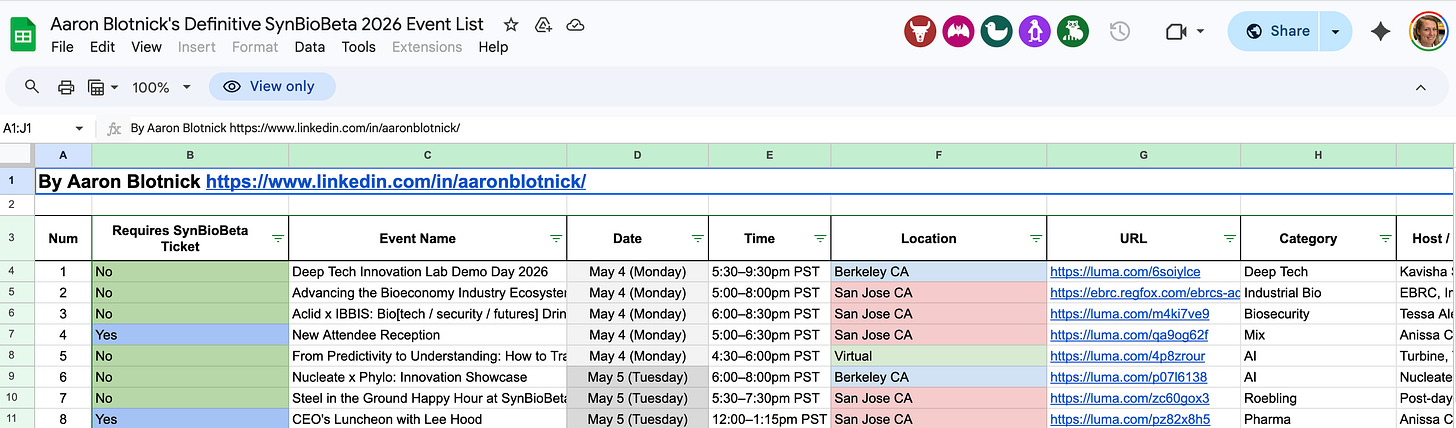
Before I leave (to go wrap up my normal work day before SynBioBeta takes me by storm in the morning), I wanted to shout out Aaron Blotnick at Bits in Bio, who curated a list of SynBioBeta-adjacent events this week:
Even if you’re not going to the conference, you can hit up many of the mixers happening in the evenings! Thanks Aaron! We haven’t met but I always appreciate you for surfacing all the good stuff!
Also if you’re reading this and aren’t on the Bits in Bio Slack, first of all, I am surprised! You will find MUCH more useful things there than here! I don’t read it every day, but it’s my #1 go-to place when I want to know about AI x bio, tech x bio, biotech events, or when someone tells me they’re looking for a job in bioinformatics, biotech, etc. It’s super active and I am definitely underutilizing it. Also if you ever come across an in-person Bits in Bio event (for example here’s one this week), I suggest going! They are among the highest quality I’ve found in the Bay Area for bio events! Always full of welcoming, smart people! Also Bits in Bio is not just for those of us in SF — they have other local chapters, and lots of virtual stuff too!
Lastly of course, come join me at SynBioBeta this week!
Ok that’s it for today! Until next time!
— Jessica